|
11/18/2023 0 Comments Peptide backbone of alpha helix
Please realize that the resulting numbers are only approximate, reflecting the sum of partial surface areas for all the dots you see. Showing dots isn't mandatory, but it's a good idea to confirm that you're getting the value for the atom dot surface you think you're using. Set dot_density, 3 remove hydro remove solvent set dot_solvent, off get_area elem N + O get_area elem C + S get_area all 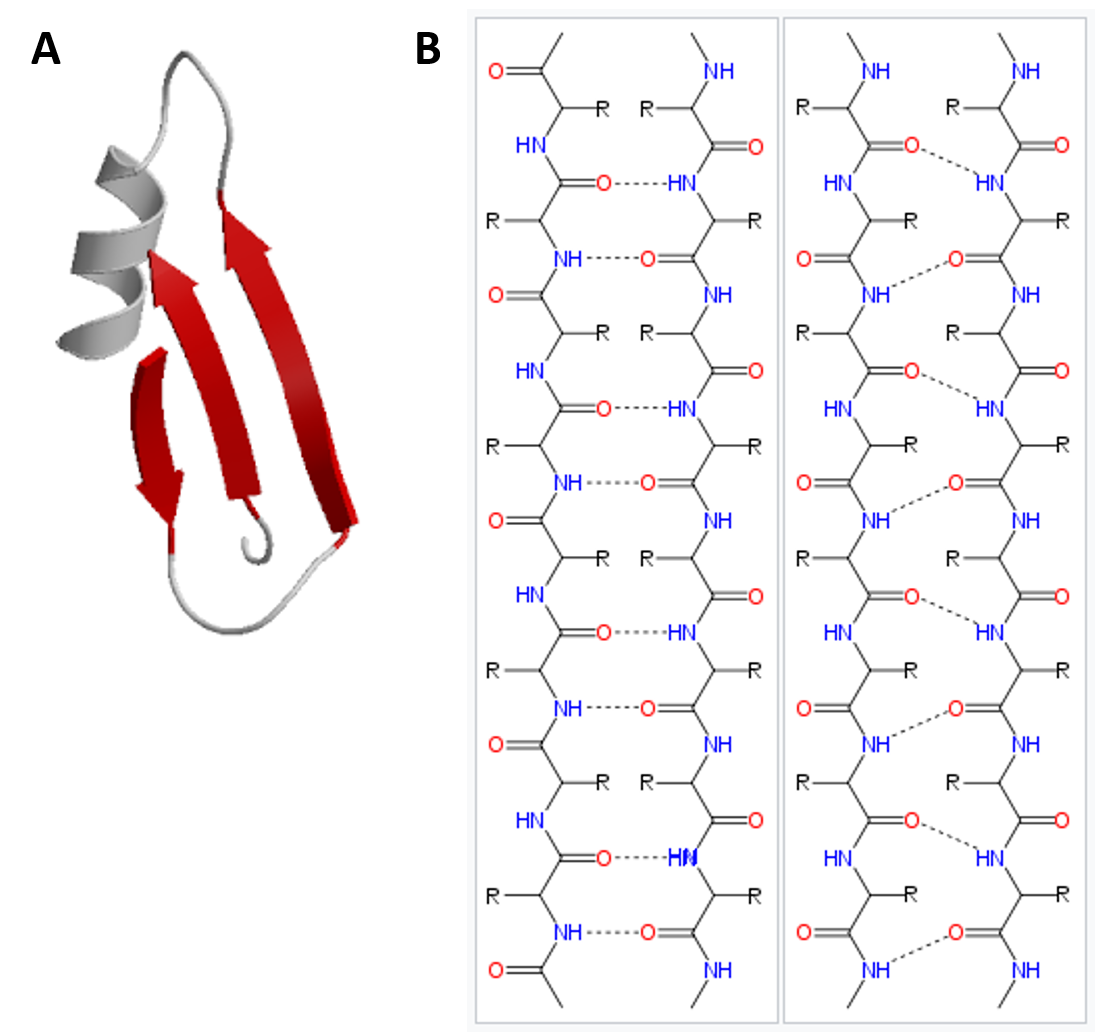 
The get_dihedral function requires four single-atom selections to work: The endpoints of the effective cutoff curve is a function of the two h_bond_cutoff_* setting. The two h_bond_power_* settings are merely fitting parameters which enable PyMOL to reproduce a curve shape reflecting Kabsch Figure 1. C-O-C), then by default, HAB cannot be less then 90 degrees. In other words, if B is only one atom (e.g. Since h_bond_cone in 180 by default, the present behavior is to simply reject any hydrogen bond where the hydrogen would lie behind the plane defined by the acceptor atom (A) in relation to its bonded atom(s) B (if any). Tthe hydrogen must also lie within a cone of space with its origin on A (along B->A) and angular width h_bond_cone. If H does not exist, then PyMOL will guess a hypothetical coordinate which may not actually be valid (in plane, etc.). With respect to the h_bond_* settings, the angle in question for h_bond_cutoff_* and h_bond_max_angle is ADH, assuming H exists. two carbonyls hydrogen bonding because they're being treated like hydroxyls). 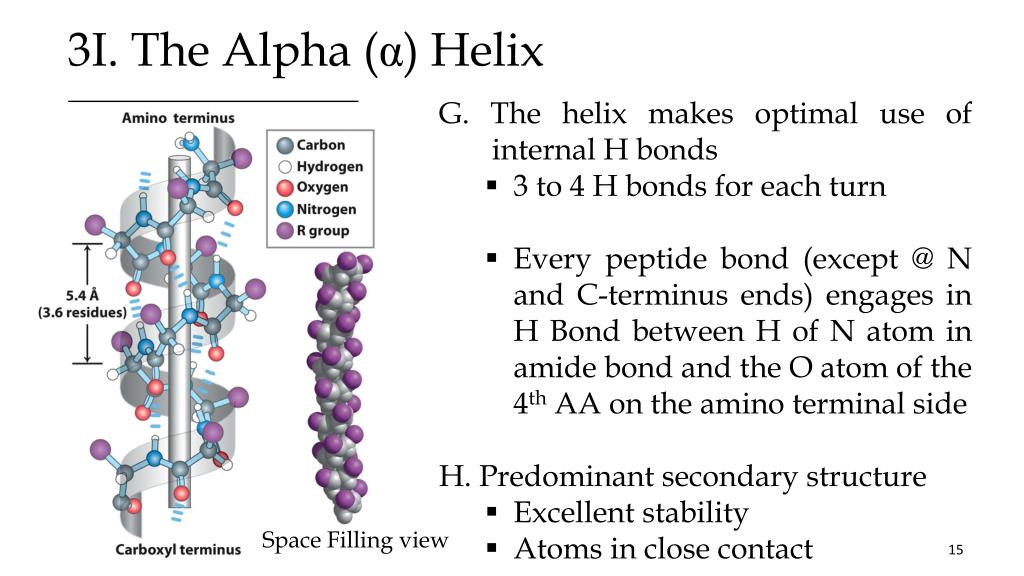
So the bottom line is that PyMOL merely offers up putative polar contacts and leaves it to the user to determine whether or not the interactions present are in fact hydrogen bonds, salt bridges, polar interactions, or merely artifacts of incorrect assignments (i.e. In addition, PyMOL would also need to consider the implied coordinate error due to overall structure resolution and local temperature factors before rigorously asserting than any specific hydrogen bond does or does not exist.įurthermore, our hydrogen bond detection machinery was originally developed for purposes of mimicking Kabsch and Sander's DSSP secondary structure assignment algorithm (Biopolymers 22, 2577, 1983) which is based on a rather generous notion of hydrogen bonding (see Kabsch Figure 1).Īlthough this approximate capability can be accessed via the distance command using mode=2, the criteria applied by our implementation may be based on heavy-atom coordinates (only) and does not necessarily correspond to anything rigorous or published. Rigorously determining the location of lone pair electrons and proton coordinates from raw PDB files is a nontrival problem especially when arbitrary small molecule structures are present. As it stands, all of those things are guessed heuristically. Generally speaking, PyMOL does not have sufficient information to rigorously determine hydrogen bonds, since typical PDB file are ambiguous with respect to charge states, bonds, bond valences, and tautomers. "Polar contacts" check geometry as well as distance. The "polar contacts" mentioned above are probably better at finding hydrogen bonds than these scripts. pdb h_add prot1 h_add prot2 select don, ( elem n, o and ( neighbor hydro )) select acc, ( elem o or ( elem n and not ( neighbor hydro ))) dist HBA, ( prot1 and acc ),( prot2 and don ), 3.2 dist HBD, ( prot1 and don ),( prot2 and acc ), 3.2 delete don delete acc hide ( hydro ) hide labels, HBA hide labels, HBD # NOTE: that you could also use this approach between two # non-overlapping selections within a single object. # EXAMPLE 2 # Show hydrogen bonds between two proteins load prot1.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
